001、生成统计文件
[root@PC1 test01]# ls ## 测试bam文件 SRR21814498.sorted.bam [root@PC1 test01]# samtools flagstat SRR21814498.sorted.bam > stat.txt ## 生成统计文件 [root@PC1 test01]# ls SRR21814498.sorted.bam stat.txt
002、结果文件
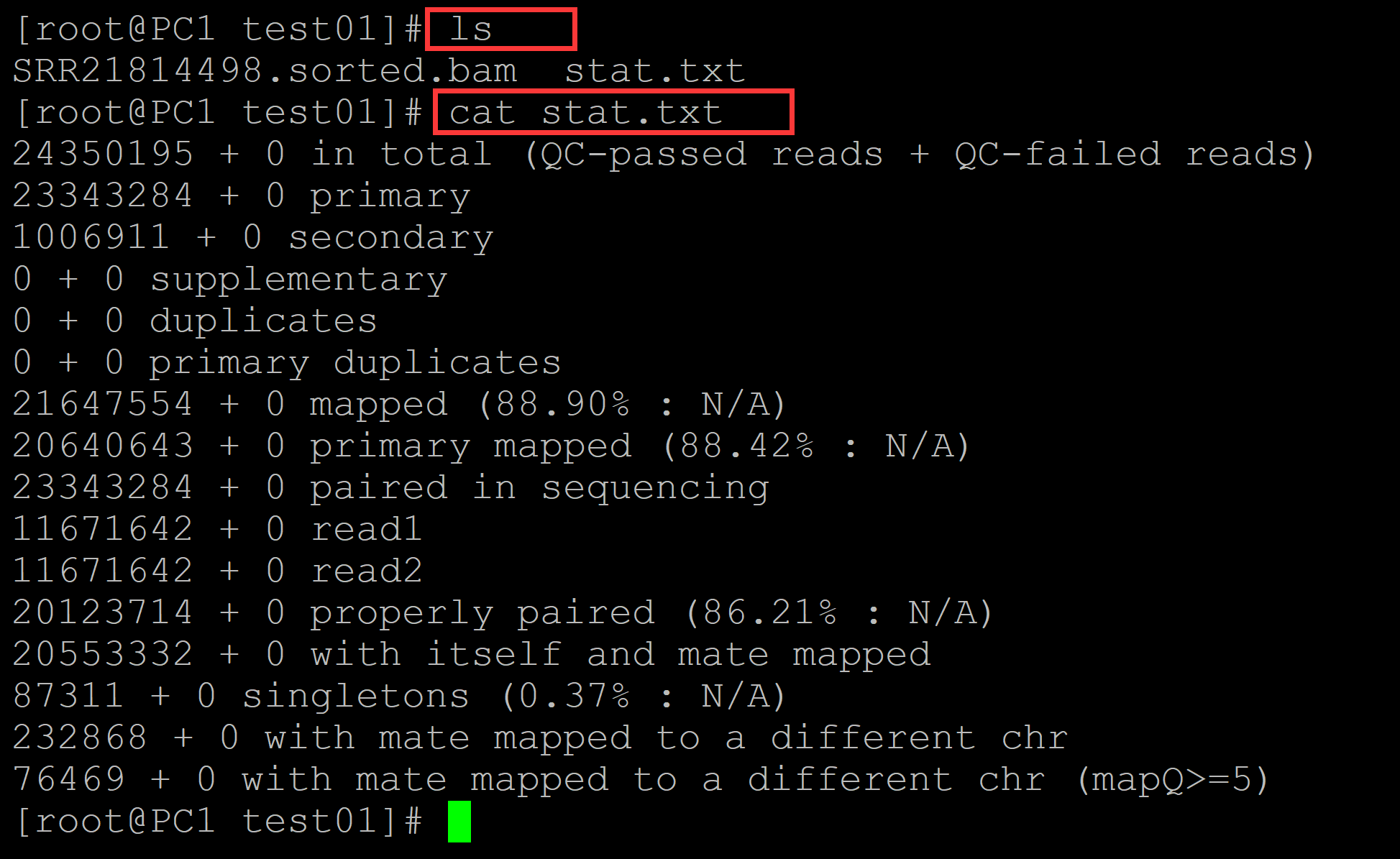
[root@PC1 test01]# cat stat.txt 24350195 + 0 in total (QC-passed reads + QC-failed reads) ## ?? 23343284 + 0 primary ## 总共的reads数目 1006911 + 0 secondary ## ??? 0 + 0 supplementary 0 + 0 duplicates 0 + 0 primary duplicates 21647554 + 0 mapped (88.90% : N/A) 20640643 + 0 primary mapped (88.42% : N/A) ## 初级匹配率 23343284 + 0 paired in sequencing 11671642 + 0 read1 ## read1中的read数目 11671642 + 0 read2 ## read2中的read数目 20123714 + 0 properly paired (86.21% : N/A) ## 完美匹配率,比对到同一条参考序列,并且两条reads之间的距离符合设置的阈值 20553332 + 0 with itself and mate mapped 87311 + 0 singletons (0.37% : N/A) 232868 + 0 with mate mapped to a different chr 76469 + 0 with mate mapped to a different chr (mapQ>=5)
。